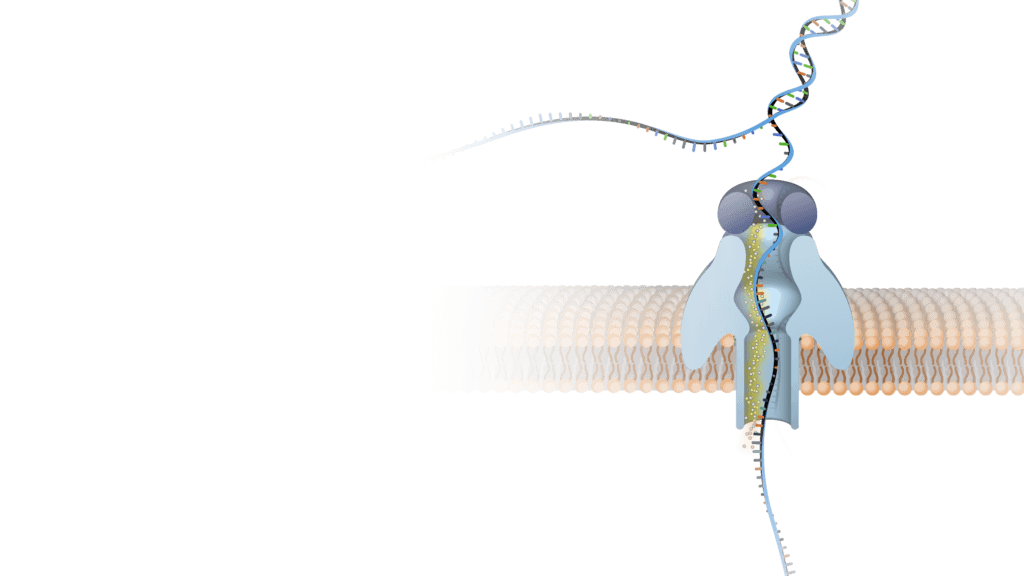
Nanopore LONG-READ sequencing technology
Whole Genome Sequencing For Any Organism at Ease
Current DNA sequencing technologies allow researchers to perform any projects, from sequencing massive genomes to accurately mapping and phasing of challenging genomic regions, such as repeats, large structural variations, and modifications.
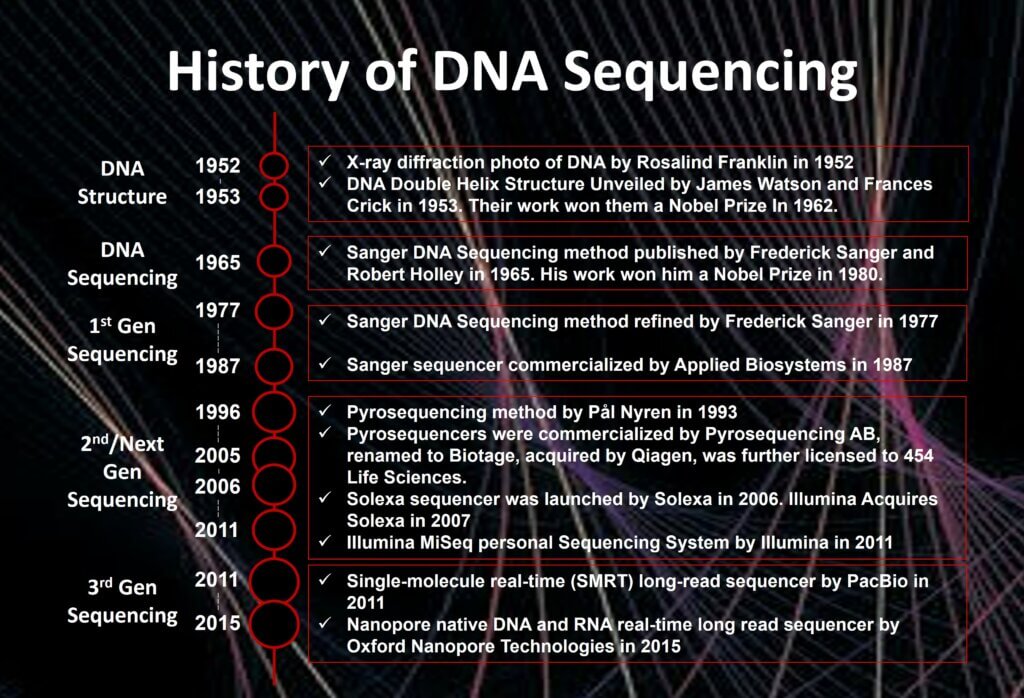
Decades of discovery science has fueled the sequencing technology revolution. As summarized in table 1, the most important accomplishments were achieved in the past 50 years by many great efforts which extended our knowledge, thinking, and capacities. For example, although it took us 13 years and cost approximately $3 billion to map a human genome, the Human Genome Project (from 1990 to 2003) has elevated our view on genomics completely, is changing the future of healthcare, and is set to continue improving health and wellbeing for patients.
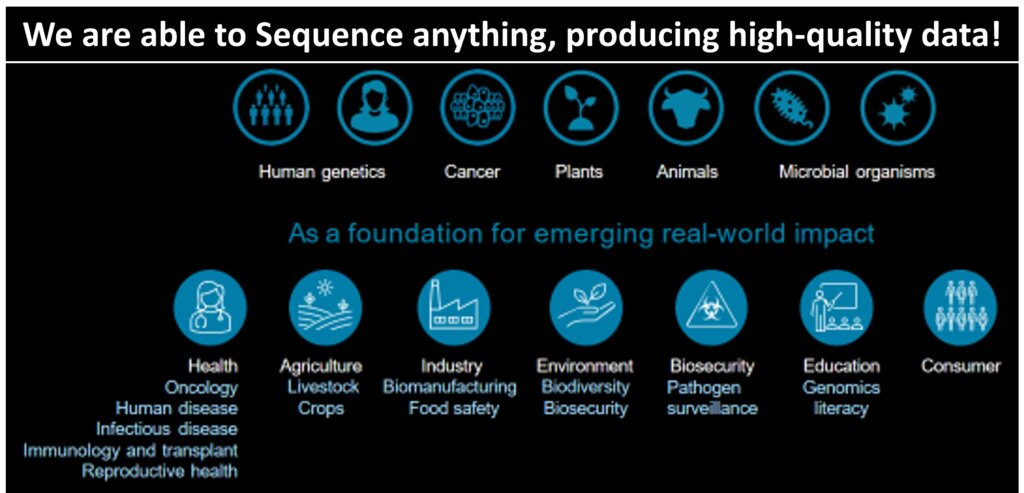
Next-generation sequencing (NGS) technologies emerged, offering faster and more cost-effective sequencing compared to traditional sequencing. The technologies have been applied to various areas, including virus genome sequencing, metagenomics, RNA sequencing, and whole genome sequencing (WGS). WGS has become a valuable tool in understanding human diseases, including cancer, and in studying various organisms (plants, animals, and microbes), and has accelerated the development of personalized medicine approaches. For example, the area of functional genomics looks at how differences in our genomes influence diseases which provides insights for developing new diagnostic methods and potential therapies, energizing precision medicine.
The field continues to evolve in improving sequencing technologies and expanding their applications. One of them is nanopore sequencing, which enables real-time analysis of native DNA or RNA molecules without involving DNA polymerases that all other technologies rely on. Nanopore sequencing is a real-time single molecule reading technology which works by monitoring the changes to an electrical current as the DNA or RNA molecule passes through a tiny protein pore, called a nanopore. This innovation led to the founding of a spin-out company, called Oxford Nanopore Technologies, in 2005.
With nanopore devices, including the portables, at anywhere on the earth, genome sequencing has never been faster or more economical than it is now. Sequencing an entire human genome can be performed in nearly three days (72 hours).
Now, we are ready to support your genomics research and are you ready for your genomic project?
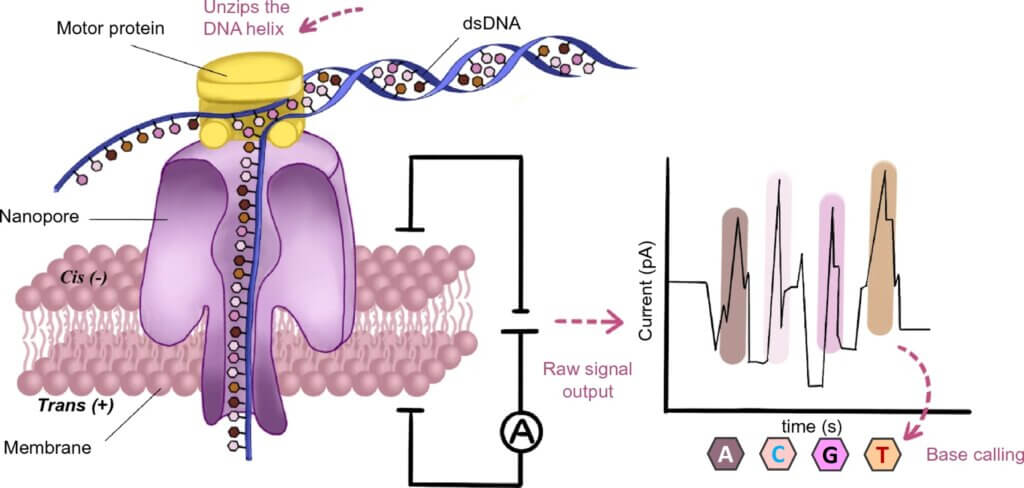
Principles of nanopore sequencing: Nanopore sequencing technology is based on the utilization of nanoscale protein pores, referred to as nanopores, as biosensors embedded within an insulating polymer membrane. By applying a constant voltage across the polymer membrane, negatively charged single-stranded DNA or RNA molecules are driven from the negatively charged side (cis) to the positively charged side (trans) of the membrane. The motor protein possesses helicase activity that unwinds the double-stranded DNA or DNA–RNA duplex into single strands that pass through the nanopore. The migration rate of the nucleic acid strands within the nanopore is also controlled by the motor protein. As the nucleic acid strand translocates through the nanopore, different bases induce different changes in the electric current. These changes are read by the signal receptor within the nanopore, and subsequent base recognition is accomplished via computational algorithms (base calling).
Principles of ONT Real-Time, Long-Read Sequencing Technology
Compared to other third-generation sequencing technologies, a key feature of nanopore sequencing is that it does not rely on DNA polymerases during the sequencing process, but directly reads DNA or RNA molecules through nanopores, achieving real-time sequencing in its true sense. Nanopore sequencing is performed by immersing a biological membrane containing nanopores in an ionic solution, applying a voltage to both sides of the phospholipid bilayer to generate an ionic current through the nanopore channels, and then driving the uncoiled single-stranded molecules to be tested through the nanopores (Fig. 1). For example, the most commonly used α-hemolysin nanopore channels with a pore size of 2.6 nm only allow the passage of single-stranded RNA or DNA. Thus, each polymer, which is an extended strand, blocks a channel when crossing the membrane, causing a change in the ionic current level. Based on the measured changes in the current level, the information of the bases can be read, and epigenetic modifications, such as DNA methylation, can be detected directly. As this process can be performed in real time, data can be generated and analyzed simultaneously.
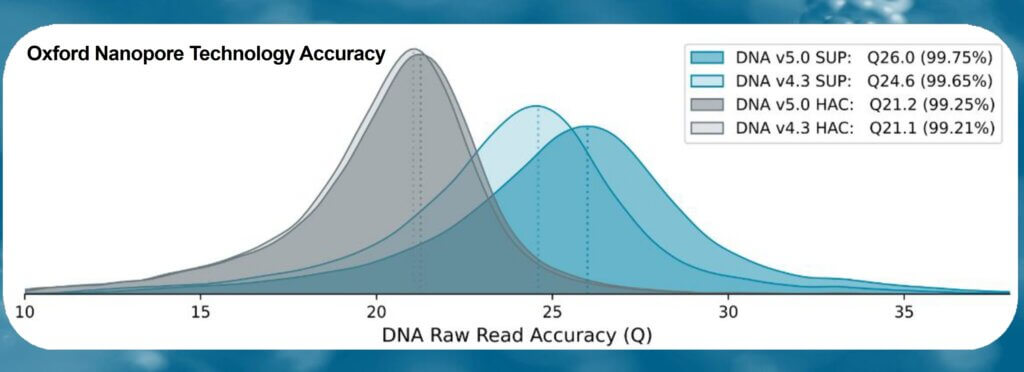
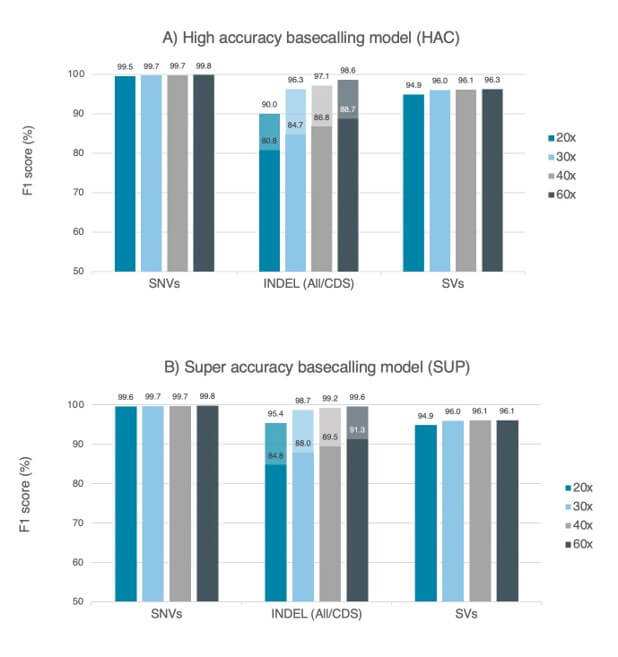
Accuracy data obtained from a dataset of 29kb N50 prepared with the Ligation Sequencing Kit V14 (with enzyme E8.2.1) and PromethION R10.4.1 Flow Cell (Oxford Nanopore Technologies). Accuracy is measured as F1 score for variant calling, using nanopore sequencing data for the human genome (HG002 cell lines) at several read depths. Variant calling was performed with wf-human-variation workflow version v2.3.0, and variants were compared against the Genome In A Bottle consortium’s HG002 truth-set (v4.2.1). SNVs and indels (< 50bp) are represented with dark colors, while indels (< 50bp) in coding regions (CDS) are displayed with lighter colors. F1 score for Dorado v0.5.2 basecalling models of (A) high accuracy (HAC, v4.3.0) and (B) super accuracy (SUP, v4.3.0).
Summary
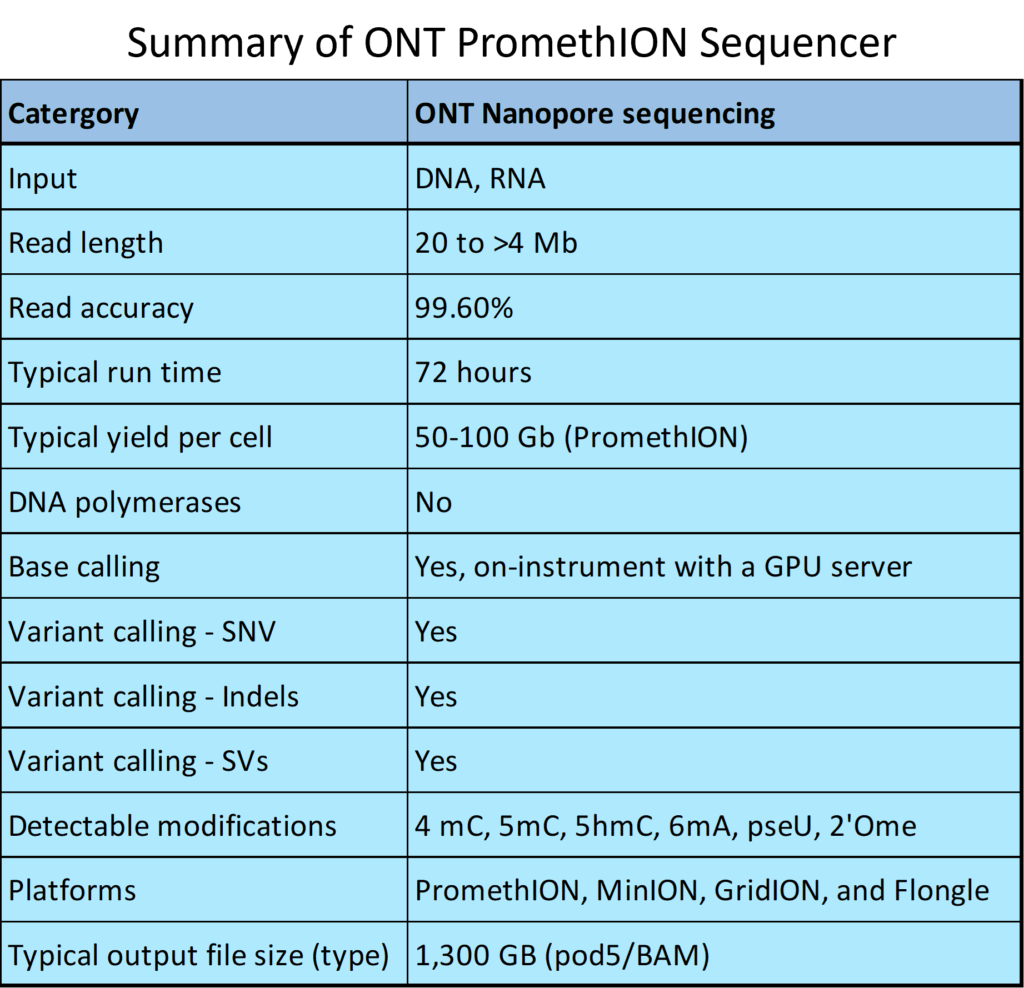
Oxford Nanopore Technologies (ONT) real-time long-read sequencing technology and library preparation kits allow to sequence native DNA or RNA, eliminating PCR bias from the data. The technology feeds a single-stranded DNA molecule through a protein nanopore and measures changes in electrical current as the molecule passes through. The single native DNA molecule long read sequencing allows for superior reading through long repetitive regions and maintains strand methylation status compared to traditional next-generation sequencing. These long reads are essential for detecting structural variants, including single nucleotide variations (SNV), structural variations (SV), insertions/deletions (Indels), copy number variants (CNV), and structural repeat regions (STR), as well as methylation modifications, and for analyzing full-length RNA transcript isoforms and poly(A) tail length.
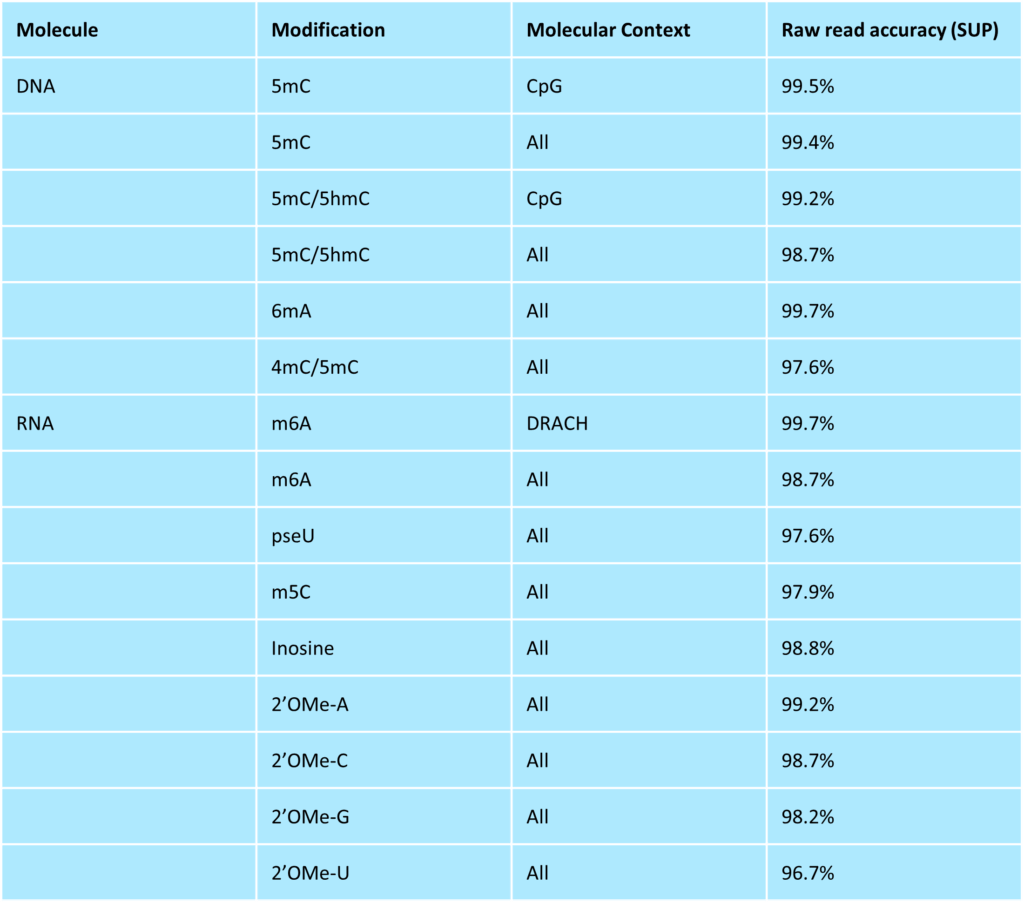
Current supported models for DNA and RNA modification basecalling available in Dorado. Accuracy values were generated on a synthetic truth-set using v5.2 SUP (for DNA) and v5.2 SUP (for RNA) basecalling models. All modification models, except RNA 2’OMe and DNA 4mC/5mC, are currently available in the latest MinKNOW version, which will be integrated in later versions.
Nanopore sequencing applications
Nanopore Whole Genome Sequencing (WGS) and Direct RNA Sequencing can be used for a variety of applications, including, but not limited to:
- The completion of existing reference genomes
- Identification of undiscovered genes and their associated function
- Characterization of uncharted regions such as centromeres and telomeres
- Identification of genomic variation (SNV, SV, Indels, CNV, STR, modified bases)
- Analyzing full-length transcripts and accurate isoform identification
- Analyzing full-length transcript and base modification
- Analyzing full-length transcript and the length of poly(A) tails
- Rapidly identifying and characterizing RNA viruses